Plants and animals have roughly 25,000 to 30,000 genes. The genes provide the information needed to make a protein, and proteins are the building blocks for all biological organisms. An ideal analogy is a blueprint (DNA) for an alternator (the protein) in a car (the plant). Proteins are the ‘parts’ for living things. Some proteins will work better than others, leading to visible differences that we call phenotypes.
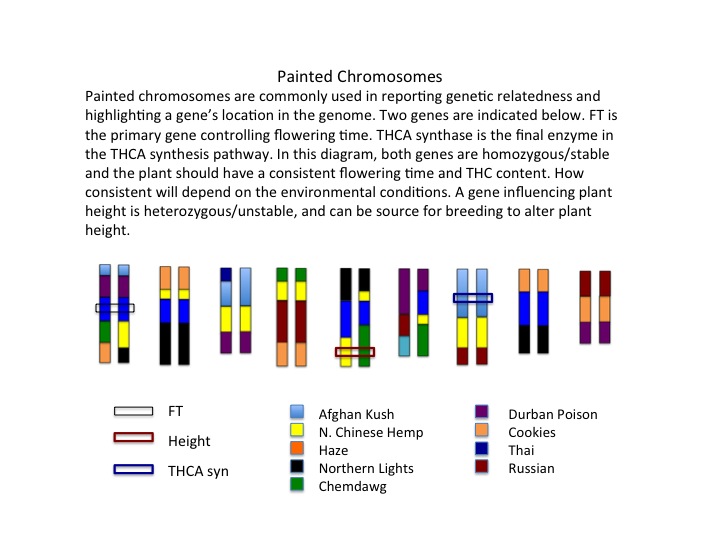 Many traits, and the genes controlling them, are of interest to the cannabis industry. For hemp seed oil, quality, quantity and content can be manipulated through breeding natural genetic variants. Hemp fibers are already some of the best in nature, due to their length and strength. Finding the genes and proteins responsible for elongating the fibers can allow for the breeding of hemp for even longer fibers. In cannabis, the two most popular genes are THCA and CBDA synthases. There are currently over 100 sequences of the THCAS/CBDAS genes, and many natural DNA variations are known. We can make a family tree using just the THCAS, gene data and identify ‘branches’ that result in high, low or intermediate THCA levels. Generally most of the DNA changes have little to no effect on the gene, but some of the changes can have profound effects.
Many traits, and the genes controlling them, are of interest to the cannabis industry. For hemp seed oil, quality, quantity and content can be manipulated through breeding natural genetic variants. Hemp fibers are already some of the best in nature, due to their length and strength. Finding the genes and proteins responsible for elongating the fibers can allow for the breeding of hemp for even longer fibers. In cannabis, the two most popular genes are THCA and CBDA synthases. There are currently over 100 sequences of the THCAS/CBDAS genes, and many natural DNA variations are known. We can make a family tree using just the THCAS, gene data and identify ‘branches’ that result in high, low or intermediate THCA levels. Generally most of the DNA changes have little to no effect on the gene, but some of the changes can have profound effects.
In fact, CBDAS and THCAS are related, in other words, they have a common ancestor. At some point the gene went through changes that resulted in the protein producing CDBA, or THCA or both. This is further supported by the fact that certain CBDAS can produce some THCA, and vice-versa. Studies into the THCAS and CBDAS family are ongoing and extensive, with terpene synthase genes following close behind.
Identifying gene (genetic) variants and characterizing their biological function allows us to combine certain genes in specific combinations to maximize yield, but determining which genes are important (gene discovery) is the first step to utilizing marker-assisted breeding.
Gene Discovery & Manipulation
The term genetics is often misused in the cannabis industry. Genetics is actually “the study of heredity and the variation of inherited characteristics.” When people say they have good genetics, what they really mean is that they have good strains, presumably with good gene variants. When people begin to cross or stabilize strains, they are performing genetic manipulation.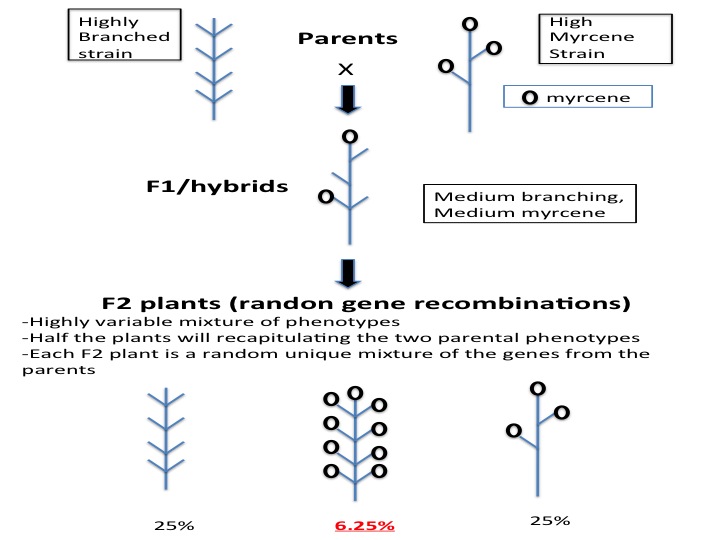
A geneticist will observe or measure two strains of interest, for example a plant branching and myrcene production. The high-myrcene plant is tall and skinny with no branching, reducing the yield. Crossing the two strains will produce F1 hybrid seeds. In some cases, F1 hybrids create unique desirable phenotypes (synergy) and the breeder’s work is completed. More often, traits act additively, thus we would expect the F1 to be of medium branching and medium myrcene production, a value between that of the values recorded for the parents (additive). Crossing F1 plants will produce an F2 population. An F2 population is comprised of the genes from both parents all mixed up. In this case we would expect the F2 progeny to have many different phenotypes. In our example, 25% of the plants would branch like parent A, and 25% of the F2 plants will have high myrcene like parent B. To get a plant with good branching and high myrcene, we predict that 6.25% (25% x 25%) of the F2 plants would have the correct combination.
The above-described scenario is how geneticists assign gene function, or generally called gene discovery. When the gene for height or branching is identified, it can now be tracked at the DNA level versus the phenotype level. In the above example, 93.5% of your F2 plants can be discarded, there is no need to grow them all to maturity and measure all of their phenotypes.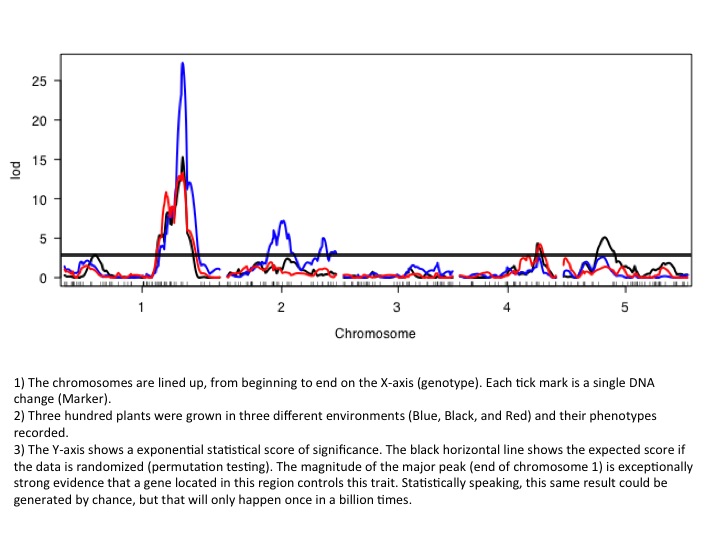
The most widely used method for gene discovery using natural genetic variation is by quantitative trait loci mapping (QTL). For these types of experiments, hundreds of plants are grown, phenotyped and genotyped and the data is statistically analyzed for correlations between genes (genotype) and traits (phenotype; figure). For example, all high-myrcene F2 plants will have one gene in common responsible for high myrcene, while all the other genes in those F2 plants will be randomly distributed, thus explaining the need for robust statistics. In this scenario, a gene conferring increased myrcene production has been discovered and can now be incorporated into an efficient marker-assisted breeding program to rapidly increase myrcene production in other desirable strains.



